Research Focus
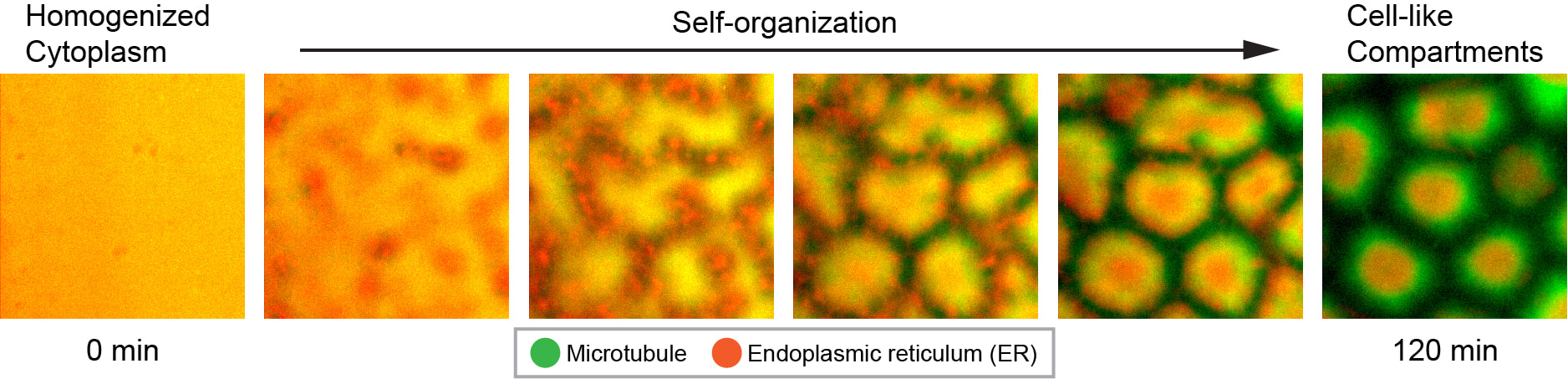
How do cytoplasmic components assemble into biologically functional spatial architecture? To answer this question, we need to observe de novo assembly of spatially organized cytoplasm. This is not possible to do in living cells as they inherit their spatial organization from mother cells instead of building it from scratch. We have recently discovered that completely scrambled frog (Xenopus laevis) egg extracts, which contain all essential components of the cytoplasm at their native concentration, are able to carry out de novo assembly of spatially organized, biologically functional cytoplasm in vitro (Read the article here). This unique property makes egg extracts an ideal model system for understanding the mechanisms of de novo cytoplasmic assembly.
Using the Xenopus egg extract system, we discovered that diffusion of protein-sized molecules (but not molecules much smaller or larger in size) becomes much faster in spatially organized cytoplasm compared to scrambled cytoplasm. Thus, self-organization could enable a broad range of biochemical reactions that would otherwise be too slow to be physiological (Read the article here). We seek to further uncover the fundamental design principles of living cellular systems by dissecting the mechanisms of self-organization in frog egg cytoplasm. These principles will be instructive to the engineering of synthetic living systems, and may shed light on the origin of cellular life.
(1) How is the microtubule (MT) and endoplasmic reticulum (ER) organization in the cytoplasm generated and maintained?
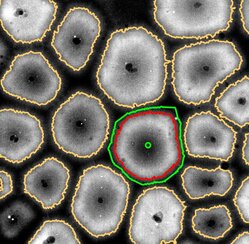
(2) What sets the intrinsic length scale of self-organized cytoplasmic patterns?
(3) How do self-organized MT and ER structures carry out cell cycle functions?
(4) How can we leverage self-organization to build synthetic cells?
How do cytoplasmic components assemble into biologically functional spatial architecture? To answer this question, we need to observe de novo assembly of spatially organized cytoplasm. This is not possible to do in living cells as they inherit their spatial organization from mother cells instead of building it from scratch. We have recently discovered that completely scrambled frog (Xenopus laevis) egg extracts, which contain all essential components of the cytoplasm at their native concentration, are able to carry out de novo assembly of spatially organized, biologically functional cytoplasm in vitro (Read the article here). This unique property makes egg extracts an ideal model system for understanding the mechanisms of de novo cytoplasmic assembly.
Using the Xenopus egg extract system, we discovered that diffusion of protein-sized molecules (but not molecules much smaller or larger in size) becomes much faster in spatially organized cytoplasm compared to scrambled cytoplasm. Thus, self-organization could enable a broad range of biochemical reactions that would otherwise be too slow to be physiological (Read the article here). We seek to further uncover the fundamental design principles of living cellular systems by dissecting the mechanisms of self-organization in frog egg cytoplasm. These principles will be instructive to the engineering of synthetic living systems, and may shed light on the origin of cellular life.
(1) How is the microtubule (MT) and endoplasmic reticulum (ER) organization in the cytoplasm generated and maintained?
(2) What sets the intrinsic length scale of self-organized cytoplasmic patterns?
(3) How do self-organized MT and ER structures carry out cell cycle functions?
(4) How can we leverage self-organization to build synthetic cells?
Approach
The lab takes a multidisciplinary approach to address the research questions. We use:
The lab takes a multidisciplinary approach to address the research questions. We use:
- Molecular biology, biochemistry, cell biology and engineering tools to probe in vitro and in vivo experimental model systems.
- in vitro: Xenopus egg extracts
- in vivo: Xenopus oocytes, eggs, cultured cells and embryos; C. elegans embryos; Sea urchin eggs and embryos.
- Time-lapse live imaging to characterize the dynamics of cytoplasmic components.
- Computational image processing to quantitatively analyze the microscopy data.
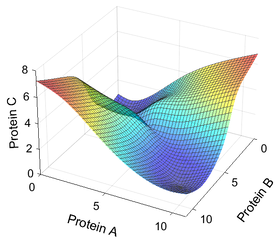
- Mathematical modeling to understand mechanisms underlying the experimental observations and to generate novel, experimentally testable predictions.